In the last few weeks (which tells you just how fast this field is evolving) the conversation in deep learning circles has shifted from using specialized fine-tuning models towards a new technology called GRAGs, short for Graphical Retrieval Augmented Generator(s). This, in turn, is forcing a rethink about the roles that knowledge graphs (and their associated portals) should play in the increasingly Wild West of hybrid knowledge systems.
The DNA of Data Representation
A DNA analogy may be appropriate here. Think of an LLM as a strand of DNA. When it comes time for an organism to replicate, the DNA splits down the middle to produce two RNA strands. In a pure model, the strand so created reflects the model completely. Such a model is usually highly optimized, is fast, and is as accurate as its source data. However, such models are also large, expensive to create, and are only accurate until the model was created, no matter how long ago that was. As most knowledge tends to remain fairly consistent, such models are called foundational - they represent a solid snapshot of knowledge at a given time that can be used for making inferences.
Carrying the analogy forward a bit, one of molecular biology's more intriguing domains is epigenetics (i.e., after genetics). With epigenetics, environmental stress factors create methyl groups that bind to the DNA at certain locations. When the DNA splits, the methyl groups cause the associated nucleic acids to appear differently to the constructed transcript RNA.
In the LLM case, this approach adds additional information to the model by “fine-tuning it”, but because this new information wasn’t part of the original model’s optimizations, it can degrade the overall optimization routines. It can increase the potential for computational hallucinations (errors that occur because the functions are no longer continuous). This approach is also stochastic in that you are essentially increasing the frequency of certain new knowledge being chosen (increasing or decreasing the weight of a given assertion) without necessarily having an indication of how that weight affects the model.
With Retrieval Augmentation Generators (RAG), on the other hand, the closer analogy would be post-facto gene splicing. In this case, the foundational model becomes one of a set of sources, where the other sources can be LLMs, PDFs, knowledge graphs, web content, or other resources. Here, the initial prompt query from the LLM is tokenized and then sent to a bridge filter that maps these tokens to an initial query result that’s then cached.
From this initial prompt, additional resources can be added (expanding upon the graph) until a specific number of iterations pass or no additional nodes are extracted from the query. Nodes can also be ranked by other arbitrary weights, such as the source's reliability or how recent the data is. Finally, the foundation LLM is queried, providing base-level data.
The final step then involves passing this data through templates to generate the appropriate output. The templating mechanism is essentially the same one that a plain LLM uses but considers the weighting of concepts as to what specifically appears in the output. The resulting concept ontology is retained locally and is used for additional inferencing and context in future queries.
What’s important here is that you’re not actually modifying the foundation graph in any way. Instead, the LLM acts purely as an aggregator, pulling together and prioritizing concepts into an appropriate response message. The immediate context learns information, but it will go out of scope with the session unless it is later captured and made part of the driving corpus.
GRAGs and Ontologies
There are several implications of GRAGs. First, a GRAG highlights the difference between a key-based ontology and a contextual one. RDF is a key-based ontology - every concept and instance within a given ontology has a specific URL that identifies that concept. In a GRAG, however, concepts are defined primarily by context, with the boundaries of such concepts being fairly fuzzy.
Both of these views of concepts have value and can actually work together. For instance, consider two related concepts - to hold and to carry. These are fairly fuzzy concepts, though one can be considered static-containing (to hold) while the other is movement-oriented (to carry). The broader category then is “to contain”. These are LLM concepts.
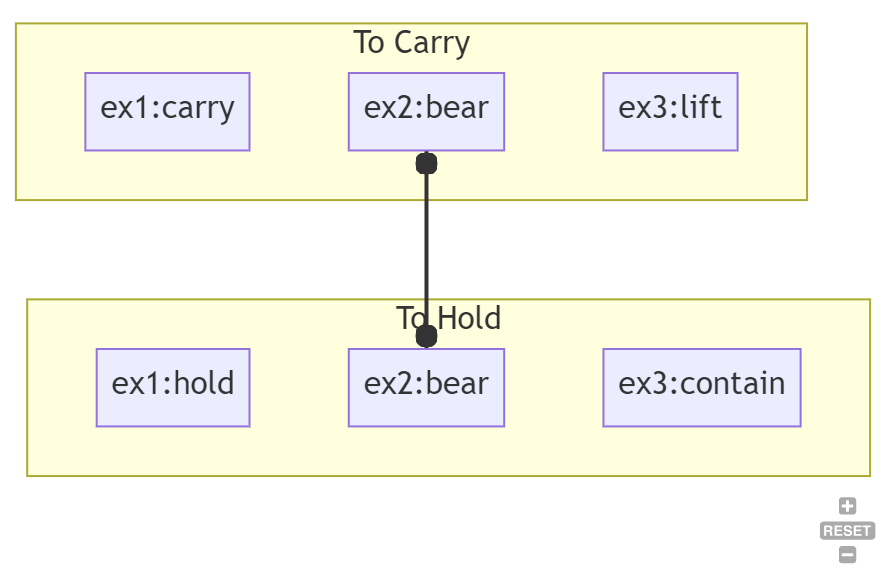
If you were to identify then six or seven different RDF concepts ( such as ex1:hold, ex1:carry, ex2:bear, ex3:contain, etc.), you could overlap the RDF concepts as points within LLM concept space. For instance, ex1:carry, ex2:bear, and ex3:lift would be in the “to carry” LLM concept, while ex1:hold, ex2:bear, and ex3:contain would be associated with the “to hold” concept.
This breakdown is important because it provides a conceptual mapping from one RDF concept to another. For instance, if you want to map the LLM concept “to carry” to ontology 1, you’d look for “to carry” terms in the ex1: namespace, i.e., ex1:carry. If you wanted to map this to ontology 3, you’d get ex3:lift. This means that you can, to a first-order approximation, create a map from ex1: to ex3: without resorting to lexical means.
You can similarly use this kind of analysis to identify ambiguous terms. For instance, ex2:bear is in the intersection between “to carry” and “to hold”. This means that in order to determine how ex2:bear is being used, you need other disambiguating information in the model (or you need some word-vec dot-product comparison to determine whether the usage of ex2:bear is closer to “to carry” or “to hold”.
Summary
I believe that the discussions about ontologies and large language models are just starting. Knowledge graphs provide a means to specify and manually manipulate local ontologies, which factor heavily in data modeling and often determine the shape of data. At the same time, no single ontology is ideal for every potential contingency. GRAGs balance the two, mixing manual and automated conceptual manipulation that strengthens both sides of the equation.
Kurt Cagle is the Editor of The Ontologist. He lives in Bellevue, Washington.